The most consistently-updated places to find all of my academic papers are my ORCID or Google Scholar pages.
Neither of those pages give me the option to explain the narrative arc of my academic career, nor to provide commentary or summaries of my favorite research projects - that’s what this page is for.
Narrative Career Arc
I did short stints in the Daunert and Van Oijen labs as an undergraduate, but spent the last couple of years of my undergraduate research career working with the late Jeff Prince, using scanning and transmission electron microscopy to answer a variety of biological questions.
Inspired by conversations with my labmates in Prince Lab, I took a short time to perform my first independent set of experiments, renting out lab space at Colorado State University which culminated in my Plant learning paper. I then went to Cambridge to work on chloroplast biotechnology with Jim Haseloff, leading to a handful of papers and an MPhil in plant biology.
I decided to return to the United States for my PhD, motivated by a frustration with the difficulty of performing field experiments using transgenic plants in the United Kingdom. Naturally, I came to the US University with the largest number of transgenic plant field experiments, University of California Davis. After a few rotations there, I chose to do my PhD with Patrick Shih. Halfway through my PhD, Patrick moved to UC Berkeley, so I left Davis with a MS and moved to the Joint BioEnergy Institute to finish the PhD under the auspices of UC Berkeley.
A bee's foot under scanning electron microscopy (SEM)

A hibiscus anther under focus-stacked light microscopy
Featured Research Projects
Plant Learning Paper
My first, and likely only, solo experimental paper. I received lots of useful advice, but only the advice I sought out - I had no direct academic supervisor, and was working out a lab space I rented myself.
This paper was an attempted replication of a big-if-true paper that came out during my senior year of undergrad. The effect didn't replicate, and the fact that nobody (including the original authors) has gotten the effect to replicate in the 4 years since suggests that it wasn't just that I didn't have the right expertise to replicate it, but rather that the phenomenon isn't real. Nonetheless, the publication process for the paper was slow and painful - journals really don't like publishing negative results!
What I learned: Experimental design and working with minimal supervision.
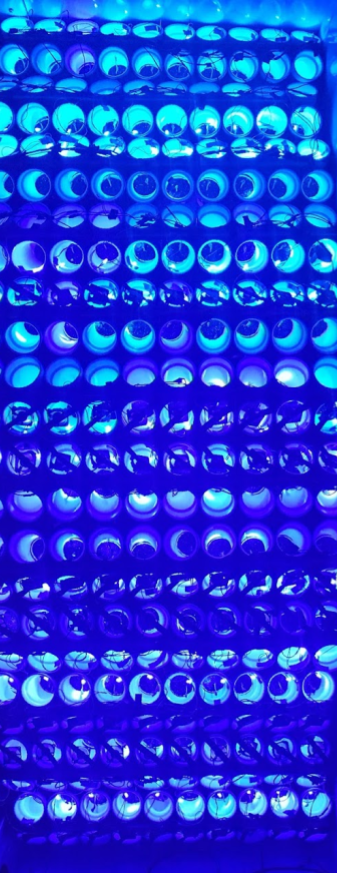

Y-shaped PVC maxes with computer fans and blue LEDs attached to pots. The lights and fans run on a schedule, and the pea plant "chooses" which tube to grow into over a several day period. They naturally grow towards light, this experiment was aiming to "train" them to grow towards the fan
Gall Paper
My gall paper was the first first-author paper of my PhD to be published, and is probably my most widely-ranging paper with respect to experimental techniques. It started with a lot of field work at UC Davis, but ultimately most of the interesting findings came from leveraging the wide range of expertise from my colleagues at JBEI.
The project taught me about working with a large team of collaborators (9 other scientists directly contributed to the experiments or data analysis, but ultimately I was responsible for orchestrating, directing, and managing the project, and either lead or assisted with most of the experiments) and interpreting data from a wide range of biological techniques.
What I learned: Collaboration, project management, and interpreting data from diverse biological techniques.
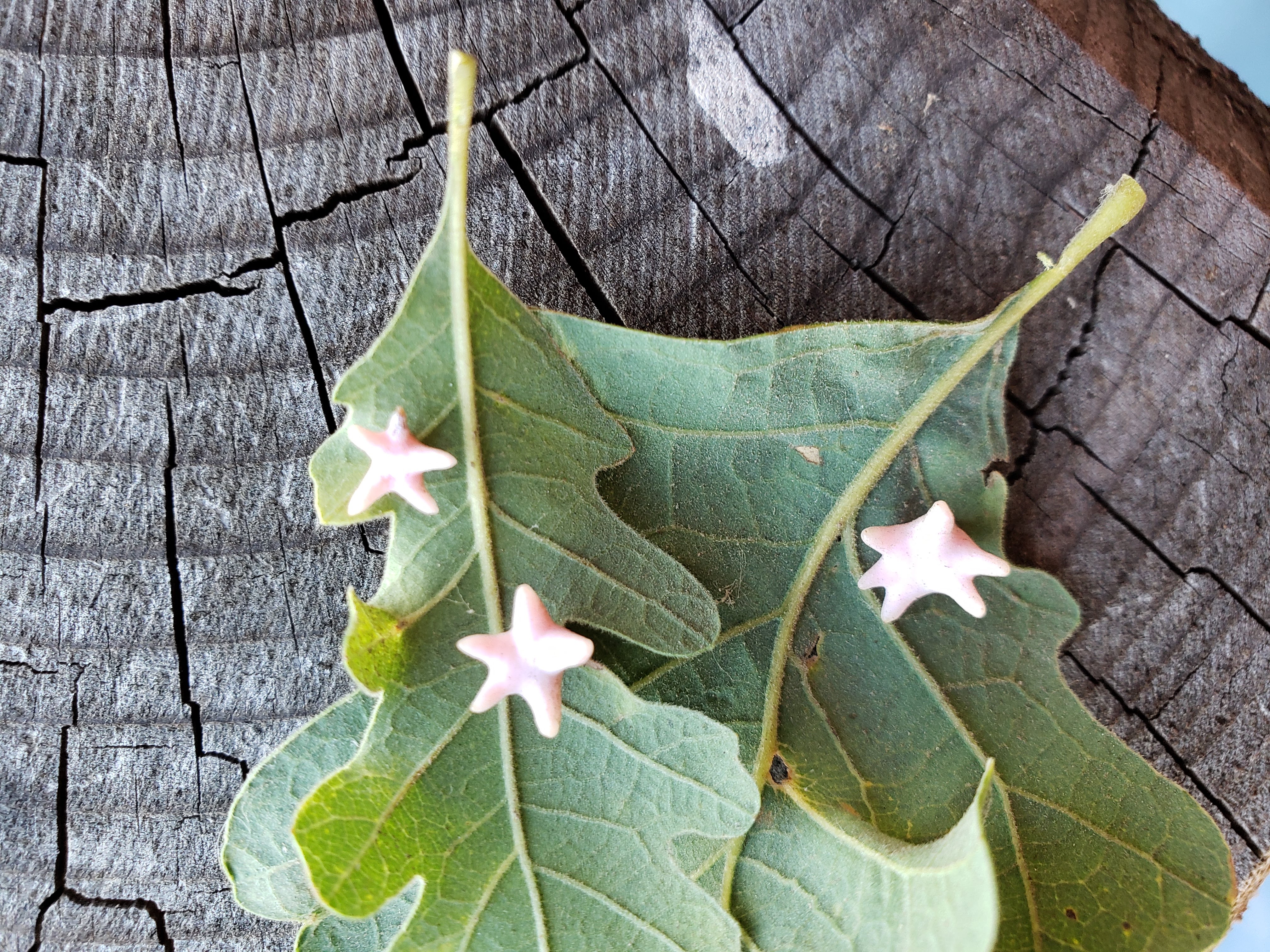

Antron douglasii galls on valley oak leaves
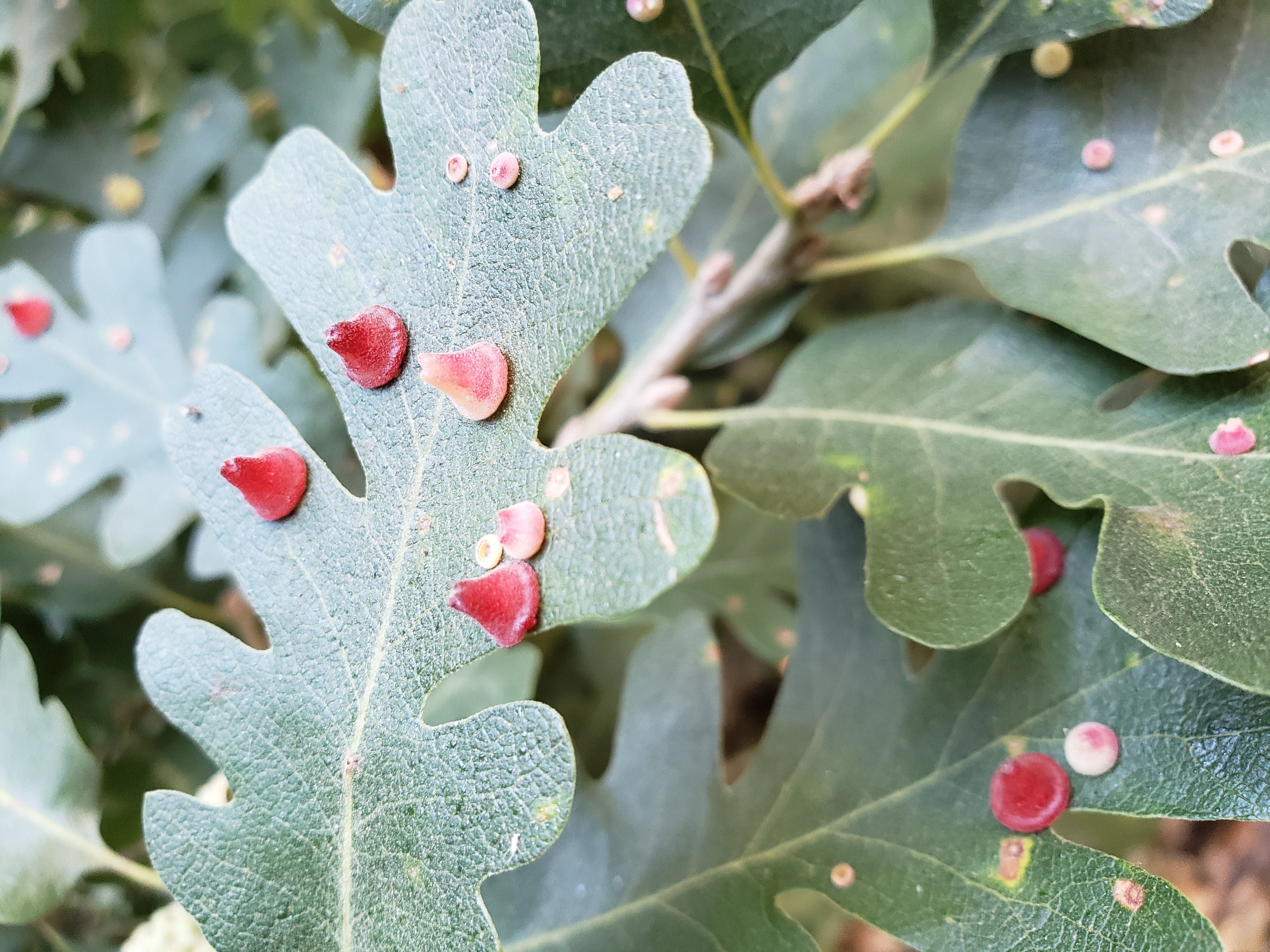

Andricus kingi galls on valley oak leaves
FTO Paper
My FTO paper was a spiritual descendant of the plant learning paper - also a replication attempt of a recent and big-if-true publication from another lab. This time, to my relief, the phenomenon replicated, which made the publication process much smoother.
This project taught me a lot about RNAseq data analysis and hypothesis generation - we confirmed that the phenomenon was real and suggest some plausible mechanisms.
What I learned: RNAseq data analysis and hypothesis generation.
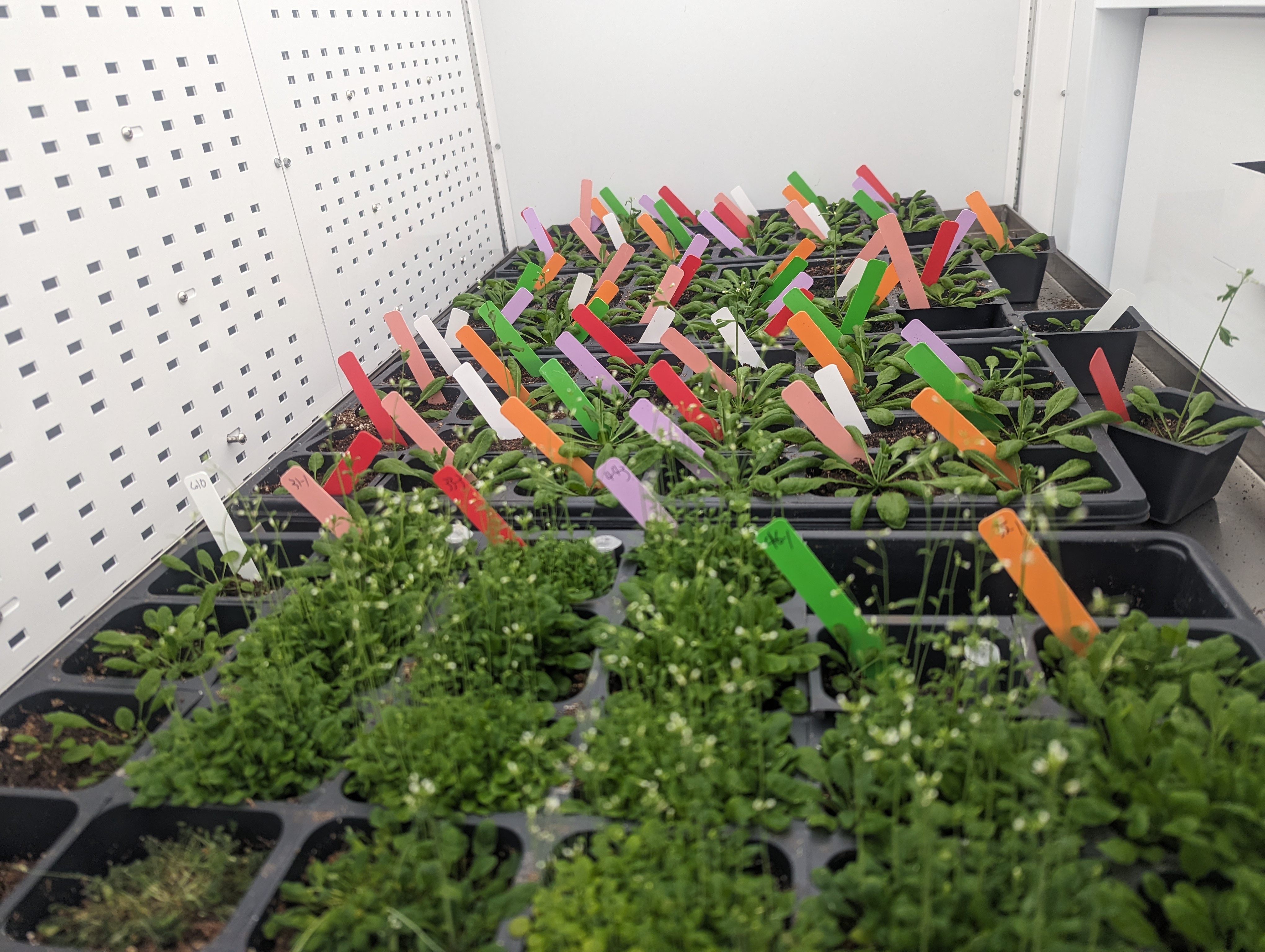

Transgenic arabidopsis plants (nursery pots in foreground, one-plant-per-pot in background) expressing a human RNA demethylase
Transcriptional Repression Toolkit Paper
My transcriptional repression paper was probably the most frustrating project I've worked on - it began as my rotation project in Shih Lab at the very start of my PhD, but was plagued with dead ends and experimental repeats, and ended up being the last of my first-author papers to be published. The paper is co-authored with two undergraduate researchers I trained and advised, both of whom are now doing PhDs of their own.
What I learned: Perseverance and management.
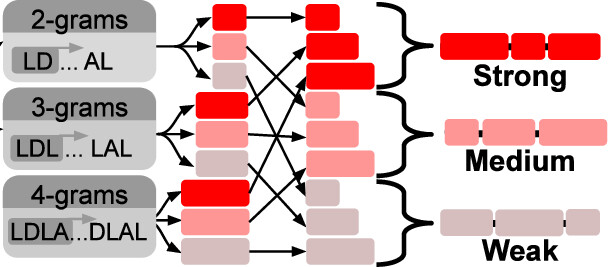
A pre-LLM "natural language processing" algorithm I applied to protein sequences right at the start of the pandemic. This is how I made synthetic transcriptional repression domains with activity predicted in advance